Projects
Research tools and applications in bioinformatics and data analysis
3 projects

dar – Differential Abundance Analysis by Consensus
An R/Bioconductor package designed to tackle robust differential abundance testing in microbiome research. Microbiome data are sparse, highly variable and compositional, making single-method approaches unstable. dar provides a consensus-based framework that integrates multiple state-of-the-art DA methods into pipeable, recipe-like workflows. Results can be aggregated using different consensus strategies like majority voting or weighted combinations, improving robustness, interpretability and reproducibility. Supports phyloseq and TreeSummarizedExperiment data structures with visualization tools and workflow export/import functionality.
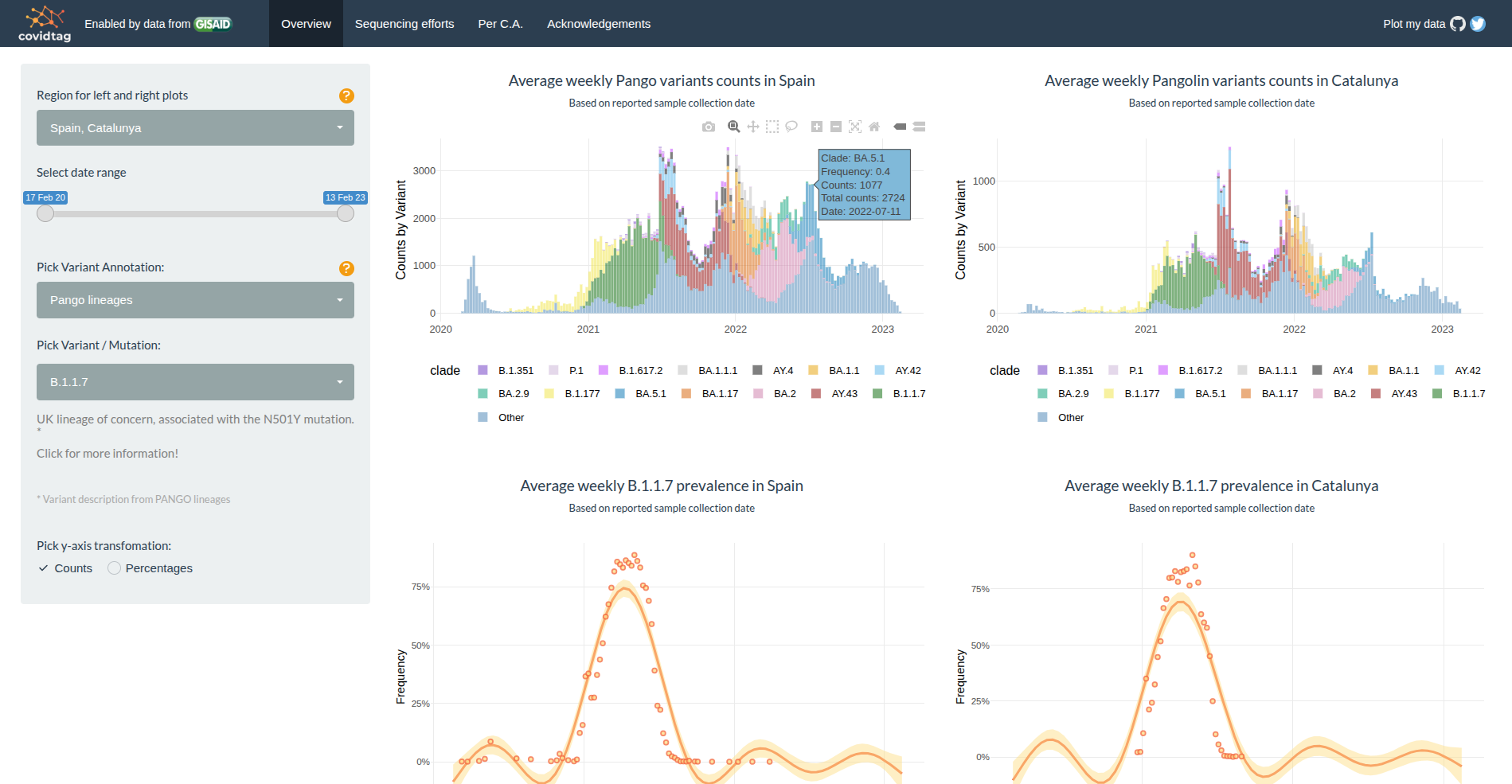
CovidTag – Tracking SARS-CoV-2 variants in Spain
A public web application designed at IrsiCaixa to monitor the spread and replacement of SARS-CoV-2 variants across Spain. The platform aggregates genomic data from GISAID and provides an up-to-date view of how viral lineages circulate in each autonomous community over time. Users can visualize variant frequencies in each region, compare communities with Spain's overall status, track how new variants displace dominant lineages, and quantify sequenced samples per territory. Weekly updates ensure the dashboard reflects the most recent genomic evidence available.

biocroxytest – Handle Long Tests in Bioconductor Packages
An R/Bioconductor package designed to work with long-running tests in Bioconductor software. Bioconductor's build system enforces strict time limits on standard test suites, making it difficult to include expensive integration tests or large-data checks. biocroxytest introduces a roxygen2-based workflow with a dedicated @longtests roclet, allowing developers to annotate long tests directly inside roxygen2 documentation comments. These annotated blocks are automatically extracted into separate test files under a longtests/ directory, enabling independent execution in specialised CI jobs without compromising daily build speed.